Predicting Experimental Evolution
The underlying causes of repeated evolution can work on several different biological levels, which make quantitative modeling challenging in many cases. Although natural selection is often seen as the dominant factor, the fixation of the fittest mutant in a population is far from certain if other adaptive mutants arise more commonly due to larger mutational target size or mutational hot spot sites. We are interested in determining how predictable evolution is, but also if we can actually make forecasts of evolution under novel conditions that have not been observed before. This requires understanding and tools to predict biological processes on different levels, which is why our research questions range from the distribution of fitness effects to the mechanistic causes of mutational hot spot sites. We are a multidisciplinary team using experiments, bioinformatics and mathematical modelling and we think a lot about questions like:
- How repeatable is evolution of the phenotypic and genetic level?
- How conserved are the genotype-to-phenotype map and mutation hot spot sites between related species?
- What is causing repeated evolution in different cases?
- Why do mutations have effects on protein function and what is the best method to predict mutational effects?
- What is the distribution of mutation rates across a gene or genome? How can we predict mutation rates?
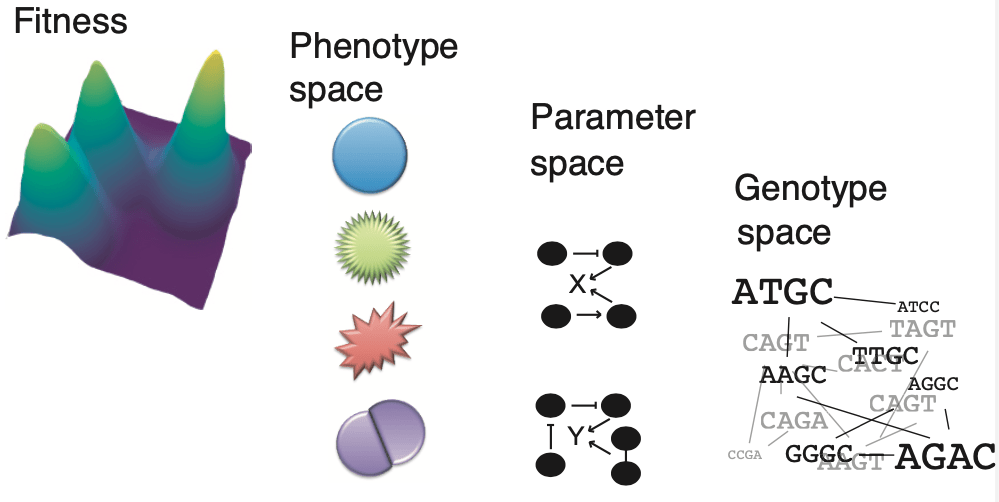
- How can we make predictive models of how mutations change molecular networks to produce adaptive phenotypes?
- Can we guide populations to a certain evolutionary trajectory?
- What is the minimal information required for successful forecasting? What needs to be measured experimentally and what can be predicted in silico?
Model Systems
Our model systems were chosen based on several considerations:
- Dominant selective pressure
- Genetic robustness and environmental robustness
- Conservation between species, but also diversity
- Knowledge of genotype-to-phenotype-to-fitness
- Intermediate complexity – several possible phenotypic and genetic solutions, regulatory molecular networks characterized
- Modularity, genotype-to-phenotype can be modified to find rare pathways
- Relevant for natural populations and medicine